Exercise
Question 1. Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine.
Sol. Nitrogenous bases are Adenine, Uracil, Cytosine, Thymine. Nucleosides (Nitrogenous base+ sugar) are Cytidine and Guanosine.
Question 2. If a double stranded DNA has 20 per cent of cytosine, calculate the per cent of adenine in the DNA.
Hint: Apply Chargaff s rule.
Sol. According to Chargafl’s rule, in a double-stranded DNA, the ratios between cytosine and guanine & adenine and thymine are constant and equals one, i.e., number of cytosine is equal to the number of guanine and the number of adenine is equal to number of thymine.
So,%C=%G
%T=%A
As per question, double-stranded DNA has 20% cytosine then according to Chargafl’s rule, it will have 20% guanine.
Total percentage of G + C content= 20% + 20% = 40% Total percentage of A+ T content= 100% -40% = 60%
Since adenine and thymine are present in equal number, content of adenine willbe30%.
Question 3. If the sequence of one strand of DNA is written as follows:
5′-AT GCATGCATGCATGCATGCATGCATGC-3′
Write down the sequence of the complementary strands in 5′ 3′ direction.
Sol. According to the complementary base pairing property of DNA, adenine present in one DNA strand always base pairs with thymine of the complementary strand and guanine in a DNA strand always base pairs with cytosine of the complementary strand and vice versa.
If the sequence ofone DNA strand is:
5′-ATGCATGCATGCATGCATGCATGCATGC-3′
Since two strands in a double-stranded DNA are antiparallel, the sequence of complementary strand in 3′ to 5′ direction will be:
3′-TACGTACGTACGTACGTACGTACGTACG-5′
The sequence of complementary strand in 5′ to 3′ direction will be: 5′-GCATGCATGCATGCATGCATGCATGCAT-3′
Question 4. If the sequence of the coding strand in a transcription unit is written as follows:
5′-AT G CAT GCA T GCA T GC AT GCATGCATGC-3′
Write down the sequence of mRNA.
Sol. Coding strand in a transcription unit is the DNA strand with polarity 5′ 3′ and does not code for any protein. It has the base sequence same as RNA except thymine at the place of uracil.
If the sequence of coding strand is:
5′-ATGCATGCATGCATGCATGCATGCATGC-3′
The sequence of mRNA will be:
5′-AUGCAUGCAUGCAUGCAUGCAUGCAUGC-3′
Question 5. Which property of DNA double helix led Watson and Crick to hypothesise a semi-conservative mode of DNA replication? Explain.
Sol. Watson and Crick observed that the nitrogenous bases are in complementary pairing in two strands of double helix of DNA molecule. Such an arrangement of DNA molecule led them to hypothesize the semiconservative mode of replication of DNA.
During DNA replication, the two DNA strands separate and act as a template for the synthesis of new complementary strands. The sequence of newly synthesized DNA is determined by the sequence of bases on the template strand based on the complementary base pairing rule. After the completion of replication, each daughter DNA molecule has one parental and one newly synthesized strand. Since only one parental strand is present from one generation to the next, this mode of replication is called semiconservative.
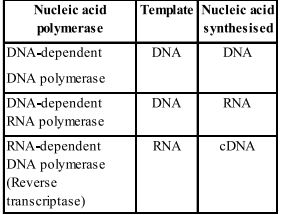
Question 6. Depending upon the chemical nature of the template (DNA or RNA) and the nature of nucleic acids synthesised from it (DNA or RNA), list the types of nucleic acid polymerases.
Sol. Different types of nucleic acid polymerases are as follows:
Question 7. How did Hershey and Chase differentiate between DNA and protein in their experiment while proving that DNA is the genetic material?
Sol. Alfred Hershey and Martha Chase (1952) worked with viruses that infect bacteria called bacteriophages. In 1952, they chose a bacteriophage known as T2 as their experimental material to prove that DNA is the genetic material. To differentiate between DNA and protein, they grew some viruses on a medium that contained radioactive phosphorus (32p) and some others on a medium that contained radioactive sulfur (35s). Viruses grown in the presence of radioactive phosphorus contained radioactive DNA but not radioactive protein because DNA contains phosphorus-containing nucleotides but protein does not. Similarly, viruses grown on radioactive sulfur contained radioactive protein but not radioactive DNA because DNA does not contain sulfur and proteins contain sulphur-containing amino acids.
Radioactive phages were allowed to attach to E. coli bacteria. Then, as the infection proceeded, the viral coats were removed from the bacteria by agitating them in a blender. The virus particles were separated from the bacteria by spinning them in a centrifuge.
Bacteria which was infected with viruses that had radioactive DNA were radioactive, indicating that DNA was the material that passed from the virus to the bacteria. Bacteria that were infected with viruses that had radioactive proteins were not radioactive. This indicates that proteins did not enter the bacteria from the viruses. DNA is therefore the genetic material that is passed from virus to bacteria.

Question 8. Differentiate between the following:
- Repetitive DNA and Satellite DNA
- mRNA and tRNA
- Template strand and Coding strand
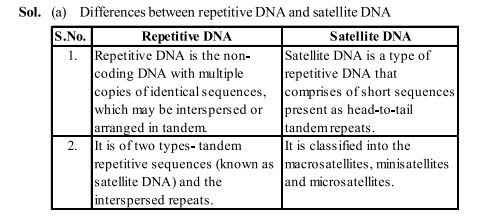
Sol. (a) Differences between repetitive DNA and satellite DNA

Question 9. List two essential roles of ribosome during translation.
Sol. A ribosome is a compact ribonucleoprotein particle which has two subunits, one large subunit and one small subunit. They associate with different mRNAs for the synthesis of different polypeptides.
Ribosome perform the following essential roles during protein synthesis or translation:
- The ribosome provides the binding site for mRNA in small subunit and two binding sites for subsequent amino acids in the large subunit and thus, be close enough to each other for the formation of a peptide bond. It also moves along mRNA by three nucleotides each time to add amino acids sequentially to the growing polypeptide. In this way, it provides an environment in which recognition between a codon of mRNA and anticodon of tRNA occurs and aminoacyl-tRNAs add amino acids to the growing chain in response to the corresponding triplet codons.
- The ribosome also acts as a catalyst. The large subunit of ribosome has peptidyl transferase activity for the formation of peptide bond between two amino acids. This activity is associated with 23S rRNA in large subunit of ribosome of bacteria so this enzyme is a ribozyme.
Question 10. In the medium where E. coli was growing, lactose was added, which induced the lac operon. Then, why does lac operon shut down some time after addition of lactose in the medium?
Sol. Lac operon is an operon in bacteria that contains structural genes responsible for metabolism of lactose. It consists of one regulatory gene, gene lac i and three structural genes, lac z, lacy and lac a (which codes for three enzymes -galactosidase, permease and transacetylase respectively). Lactose is the substrate for the enzyme -galactosidase and it regulates switching on and off of the operon. Hence, it is called inducer. In the absence of a preferred carbon source such as glucose, if lactose is provided in the growth medium of the bacteria, the lactose is transported into the cells through the action of permease. Lactose binds to the repressor and prevents its binding to operator. This allows RNA polymerase access to the promoter and transcription of structural genes proceeds leading to the formation of galactosidase. This enzyme breaks down the lactose into glucose and galactose.
After some time, lac operon shuts down due to the following reasons:
- As inducer is completely metabolized, lactose is not available to inactivate the repressor. This results in binding of repressor to the operator gene and in turn, preventing RNA polymerase from transcribing the operon. Hence, the transcription is stopped and the operon is switched off.
- Lac operon cannot be switched on by lactose due to the accumulation of glucose which is a preferred energy source over lactose.
Question 11. Explain (in one or two lines) the function of the following:
(a) Promoter (b) tRNA (c) Exons
Sol. (a) Promoter: Promoter is one of the three regions of transcription unit located towards 5′ end of the structural gene. It is a DNA sequence that provides binding site for RNA polymerase to initiate transcription. Also, the position of a promoter in a transcription unit defines the template and coding strands.
tRNA: tRNA or transfer RNA is a 75 to 85 long ribonucleic acid folded to form a secondary structure shaped like a cloverleaf Each tRNA molecule has an anticodon loop that has bases complementary to the code, and an amino acid acceptor end to which it binds to amino acids. Each tRNA has only one type of anticodon and attaches to a specific amino acid. Its function is to bind to an activated amino acid, transfer it from cellular pool to ribosomes where tRNA binds to correct codon in mRNA and helps assemble amino acids into polypeptides.
Exons: Exons are coding sequences interrupted by non-coding sequences (called introns) in the split genes of eukaryotes. Introns are removed and exons are joined to produce functional RNA by splicing. The exons in functional mRNA are transcribed into functional polypeptide.
Question 12. Why is the Human genome project called a mega project?
Sol. Human genome project aimed to sequence every base in human genome. It was a 13-year project coordinated by the U.S. Department of Energy and the National Institute of Health. The project was completed in 2003.
It was a mega project due to the following reasons:
- Large size of human genome and high invested cost of sequencing: Human genome is said to have approximately 3 x 109 bp and the total estimated cost of the project would be approximately 9 billion US dollars, and if the cost of sequencing required is US$ 3 per bp (the estimated cost in the beginning).
- Requirement of high-speed computational devices for data storage: The enormous amount of data would be expected to generate from DNA sequencing from a single human cell. This necessitated the use of high speed computational devices for data storage and retrieval, and analysis. HGP was closely associated with the rapid development of a new area in biology called bioinformatics.
Question 13. What is DNA fingerprinting? Mention its application.
Sol. • DNA fingerprinting is a technique used to identify individuals based on the differences in some specific regions in DNA sequence called as repetitive DNA. It works on the principle of polymorphism in DNA sequences. This technique was initially developed by Sir Alec Jeffreys who discovered that certain regions of DNA showed variations in the number of tandem repeats known as variable number of tandem repeats (VNTRs). He used a satellite DNA as probe that shows very high degree of polymorphism.
- Application of DNA fingerprinting:
- It can be applied in forensic science to identify an individual in criminal and civil cases. DNA fingerprinting is the basis of paternity testing, in case of disputes. It is used to identify genes connected with hereditary diseases. It is used to identify and protect the commercial varieties of crops and livestock.
- It finds its application in evolutionary biology where it is used to find out the evolutionary history of an organism and trace out the linkages between groups of various organisms.
Question 14. Briefly describe the following:
- Transcription
- Polymorphism
- Translation
- Bioinformatics
Sol. (a) Transcription
- It is the process in which genetic information from DNA is copied into RNA.
- In transcription, only a segment of DNA is copied into RNA. The segment of DNA that codes for RNA is called transcription unit. A transcription unit in DNA has three regions namely promoter, structural gene and a terminator. The promoter and terminator flank the structural gene in a transcription unit. The promoter is a DNA sequence that provides binding site for RNA polymerase. The terminator defines the end of the process of transcription. Also during transcription, one of the strands of DNA acts a template to direct the synthesis of complementary RNA.
- RNA is transcribed from DNA strand with polarity 3′ 5′ by the enzyme DNA-dependent RNA polymerase in 5′ 3′ direction using ribonucleotides.
- The process of transcription completes in three steps: initiation, elongation and termination. During initiation, RNA polymerase binds to promoter and initiates transcription (initiation). It uses nucleoside triphosphates as substrate.
(b) Polymorphism
- Polymorphism refers to the variation at genetic level arising due to mutations.
- DNA polymorphism is an inheritable mutation observed in a population at high frequency. The probability of such variation is higher in non-coding DNA sequence as mutations in these sequences may not have any immediate effect or impact in an individual’s reproductive ability.
- These mutations keep on accumulating generation after generation, and form one of the bases of variability or polymorphism. There is a variety of different types of polymorphisms ranging from single nucleotide change to very large scale changes. For evolution and speciation, such polymorphisms play very important role.
- Polymorphism in DNA sequence also forms the basis of genetic mapping of human genome as well as of DNA fingerprinting.
- Translation
- Translation is the process of protein synthesis from mRNA. The mRNA provides the template, tRNA brings amino acids and reads the genetic code, and rRNAs play structural and catalytic role during translation.
- During translation, amino acids are polymerised to form a polypeptide. The order and sequence of amino acids in a polypeptide are defined by the sequence of bases in the mRNA.
- The steps in translation are:
- The ribosome binds to mRNA at a specific area.
- The ribosome starts matching tRNA anticodon sequences to the mRNA codon sequence.
- Each time a new tRNA comes into the ribosome, the amino acid that it was carrying gets added to the elongating polypeptide chain. The ribosome continues until it hits a stop sequence, then it releases the polypeptide and the mRNA.
- The polypeptide forms into its native shape and starts acting as a functional protein in the cell.
Note: In bacteria, transcription and translation take place in the same compartment and many times the translation can begin much before the mRNA is fully transcribed because there is no separation of cytosol from nucleus in bacteria. In contrast, in eukaryotes, transcription takes place in nucleus and translation occurs in cytoplasm.
- Bioinformatics
- Bio informaticsis an inter disciplinary field mainly involving molecular biology and genetics, computer science, mathematics, and statistics. This field develops methods and software tools for understanding biological data. It helps to analyse and compare protein sequences, search for genes, assemble genomes, compare species based on their genetic material, forecast protein structures and find active parts of proteins, analyse gene expression, and inverse engineer genetic networks.
- It also helps to relate the genome to its phenotype and can trace evolution back to its roots to find adaptations and changes in the organisms.
Related Articles:
